组蛋白乳酸化修饰在肝纤维化中的调控作用
DOI: 10.12449/JCH260328
-
摘要: 肝纤维化是多种慢性肝病共有的可逆性病理过程,与肝硬化和肝癌等严重肝脏疾病的发生发展密切相关,已成为全球范围内的重大健康问题之一。近年来,新发现的表观遗传修饰组蛋白乳酸化被证实参与调控肝纤维化进程。本文系统综述了组蛋白乳酸化在炎症微环境与肝纤维化相互作用中的核心调控作用,旨在阐明“炎症-肝纤维化”级联调控机制,为肝纤维化的早期诊断、靶向干预及阻断恶性转化提供新视角。Abstract: Hepatic fibrosis is a reversible pathological process in various chronic liver diseases and is closely associated with the development and progression of severe liver diseases such as liver cirrhosis and hepatocellular carcinoma, and it has emerged as a significant global health challenge. In recent years, studies have shown that histone lactylation, a newly discovered epigenetic modification, actively participates in regulating the progression of hepatic fibrosis. This article systematically reviews the core regulatory effect of histone lactylation modification in the interaction between inflammatory microenvironment and hepatic fibrosis, in order to clarify the cascade regulatory mechanism of “inflammation-hepatic fibrosis” and provide new insights for early diagnosis, targeted intervention, and prevention of malignant transformation in hepatic fibrosis.
-
Key words:
- Histone Lactylation /
- Hepatic Fibrosis /
- Hepatic Stellate Cells
-
注: a,肝星状细胞的代谢重编程驱动组蛋白乳酸化修饰;b,组蛋白乳酸化修饰的表观遗传调控结果。HSC,肝星状细胞;Smad,母系抗十五表态蛋白同源物;TGF-β,转化生长因子β;TNF-α,肿瘤坏死因子α;IL-6,白细胞介素6;α-SMA,α-平滑肌肌动蛋白;COL1A1,Ⅰ型胶原α1;TIMP1,基质金属蛋白酶抑制剂1;HK2,己糖激酶2;LDH,乳酸脱氢酶。
图 1 组蛋白乳酸化修饰调控肝纤维化进程的模式图
Figure 1. Schematic diagram of histone lactylation modification regulating hepatic fibrosis progression
表 1 靶向组蛋白乳酸化抗肝纤维化策略
Table 1. Targeting histone lactylation: Therapeutic strategies against hepatic fibrosis
靶点 干预手段 模型体系 效应 文献依据 HK2 条件性敲除HK2 肝纤维化小鼠
模型乳酸水平↓、H3K18la↓、α-SMA/COL1A1↓ [29] LDHA/LDHB siRNA敲低+shRNA腺病
毒抑制人LX-2细胞/小鼠
模型H3K18la↓、HSC迁移/增殖↓、糖酵解↓ [29,31] IGF2BP2 CRISPR-Cas9敲除 HSC细胞系 ALDOA↓、乳酸生成↓、HSC活化标志物↓ [30] 乳酸化竞争 HDAC抑制剂(Apicidin/
MS275)活化HSC 乙酰化↑、乳酸化↓、纤维化基因表达↓ [10] 乳酰辅酶A合成 ACSS2抑制剂(如靶向化
合物)体外HSC模型
(待验证)理论:阻断乳酸化底物供应 [8](机制推测) MCT1转运体 AZD3965(MCT1抑制剂) 巨噬细胞-HSC共
培养核内乳酸↓、H3K18la↓、TGF-β释放↓
(预测)[27](跨研究
引申)注:HK2,己糖激酶2;H3K18la,组蛋白H3第18位赖氨酸乳酸化;α-SMA,α-平滑肌肌动蛋白;COL1A1,Ⅰ型胶原α1;LDHA/LDHB,乳酸脱氢酶A/乳酸脱氢酶B;IGF2BP2,胰岛素样生长因子2 mRNA结合蛋白2;ALDOA,醛缩酶A;HSC,肝星状细胞;HDAC,组蛋白脱乙酰酶;ACSS2,乙酰辅酶A合成酶2;MCT1,单羧酸转运蛋白1;TGF-β,转化生长因子β。↑表示上调,↓表示下调。
-
[1] YANG X, LI Q, LIU WT, et al. Mesenchymal stromal cells in hepatic fibrosis/cirrhosis: From pathogenesis to treatment[J]. Cell Mol Immunol, 2023, 20( 6): 583- 599. DOI: 10.1038/s41423-023-00983-5. [2] LIAO ZH, XIE ZY. Research progress in molecular mechanism of hepatic fibrosis and related therapeutic targets[J]. J Jilin Univ Med Ed, 2024, 50( 5): 1450- 1456. DOI: 10.13481/j.1671-587X.20240532.廖昭辉, 谢正元. 肝纤维化发病的分子机制及其相关治疗靶点的研究进展[J]. 吉林大学学报(医学版), 2024, 50( 5): 1450- 1456. DOI: 10.13481/j.1671-587X.20240532. [3] ZHONG L, ZHAO JQ, HUANG L, et al. Runx2 activates hepatic stellate cells to promote liver fibrosis via transcriptionally regulating Itgav expression[J]. Clin Transl Med, 2023, 13( 7): e1316. DOI: 10.1002/ctm2.1316. [4] TSUCHIDA T, FRIEDMAN SL. Mechanisms of hepatic stellate cell activation[J]. Nat Rev Gastroenterol Hepatol, 2017, 14( 7): 397- 411. DOI: 10.1038/nrgastro.2017.38. [5] LV XY, LV YY, DAI XF. Lactate, histone lactylation and cancer hallmarks[J]. Expert Rev Mol Med, 2023, 25: e7. DOI: 10.1017/erm.2022.42. [6] KOPPENOL WH, BOUNDS PL, DANG CV. Otto Warburg’s contributions to current concepts of cancer metabolism[J]. Nat Rev Cancer, 2011, 11( 5): 325- 337. DOI: 10.1038/nrc3038. [7] GAO X, PANG CY, FAN ZY, et al. Regulation of newly identified lysine lactylation in cancer[J]. Cancer Lett, 2024, 587: 216680. DOI: 10.1016/j.canlet.2024.216680. [8] ZHU RX, YE XL, LU XT, et al. ACSS2 acts as a lactyl-CoA synthetase and couples KAT2A to function as a lactyltransferase for histone lactylation and tumor immune evasion[J]. Cell Metab, 2025, 37( 2): 361- 376. DOI: 10.1016/j.cmet.2024.10.015. [9] LEGUBE G, TROUCHE D. Regulating histone acetyltransferases and deacetylases[J]. EMBO Rep, 2003, 4( 10): 944- 947. DOI: 10.1038/sj.embor.embor941. [10] MORENO-YRUELA C, ZHANG D, WEI W, et al. Class I histone deacetylases(HDAC1-3) are histone lysine delactylases[J]. Sci Adv, 2022, 8( 3): eabi6696. DOI: 10.1126/sciadv.abi6696. [11] KADOCH C, CRABTREE GR. Mammalian SWI/SNF chromatin remodeling complexes and cancer: Mechanistic insights gained from human genomics[J]. Sci Adv, 2015, 1( 5): e1500447. DOI: 10.1126/sciadv.1500447. [12] HU XL, HUANG XW, YANG Y, et al. Dux activates metabolism-lactylation-MET network during early iPSC reprogramming with Brg1 as the histone lactylation reader[J]. Nucleic Acids Res, 2024, 52( 10): 5529- 5548. DOI: 10.1093/nar/gkae183. [13] ZHAI GJ, NIU ZP, JIANG ZX, et al. DPF2 reads histone lactylation to drive transcription and tumorigenesis[J]. Proc Natl Acad Sci USA, 2024, 121( 50): e2421496121. DOI: 10.1073/pnas.2421496121. [14] NUÑEZ R, SIDLOWSKI PFW, STEEN EA, et al. The TRIM33 bromodomain recognizes histone lysine lactylation[J]. ACS Chem Biol, 2024, 19( 12): 2418- 2428. DOI: 10.1021/acschembio.4c00248. [15] LIU RL, REN XL, PARK YE, et al. Nuclear GTPSCS functions as a lactyl-CoA synthetase to promote histone lactylation and gliomagenesis[J]. Cell Metab, 2025, 37( 2): 377- 394. DOI: 10.1016/j.cmet.2024.11.005. [16] GALLE E, WONG CW, GHOSH A, et al. H3K18 lactylation marks tissue-specific active enhancers[J]. Genome Biol, 2022, 23( 1): 207. DOI: 10.1186/s13059-022-02775-y. [17] CERTO M, TSAI CH, PUCINO V, et al. Lactate modulation of immune responses in inflammatory versus tumour microenvironments[J]. Nat Rev Immunol, 2021, 21( 3): 151- 161. DOI: 10.1038/s41577-020-0406-2. [18] HANAHAN D, WEINBERG RA. Hallmarks of cancer: The next generation[J]. Cell, 2011, 144( 5): 646- 674. DOI: 10.1016/j.cell.2011.02.013. [19] MURRAY PJ, WYNN TA. Protective and pathogenic functions of macrophage subsets[J]. Nat Rev Immunol, 2011, 11( 11): 723- 737. DOI: 10.1038/nri3073. [20] WYNN TA, CHAWLA A, POLLARD JW. Macrophage biology in development, homeostasis and disease[J]. Nature, 2013, 496( 7446): 445- 455. DOI: 10.1038/nature12034. [21] IRIZARRY-CARO RA, MCDANIEL MM, OVERCAST GR, et al. TLR signaling adapter BCAP regulates inflammatory to reparatory macrophage transition by promoting histone lactylation[J]. Proc Natl Acad Sci USA, 2020, 117( 48): 30628- 30638. DOI: 10.1073/pnas.2009778117. [22] DICHTL S, LINDENTHAL L, ZEITLER L, et al. Lactate and IL6 define separable paths of inflammatory metabolic adaptation[J]. Sci Adv, 2021, 7( 26): eabg3505. DOI: 10.1126/sciadv.abg3505. [23] ZHANG D, TANG ZY, HUANG H, et al. Metabolic regulation of gene expression by histone lactylation[J]. Nature, 2019, 574( 7779): 575- 580. DOI: 10.1038/s41586-019-1678-1. [24] MA W, AO SX, ZHOU JP, et al. Methylsulfonylmethane protects against lethal dose MRSA-induced sepsis through promoting M2 macrophage polarization[J]. Mol Immunol, 2022, 146: 69- 77. DOI: 10.1016/j.molimm.2022.04.001. [25] GU J, ZHOU JR, CHEN QY, et al. Tumor metabolite lactate promotes tumorigenesis by modulating MOESIN lactylation and enhancing TGF-β signaling in regulatory T cells[J]. Cell Rep, 2022, 39( 12): 110986. DOI: 10.1016/j.celrep.2022.110986. [26] LI LN, LI WW, XIAO LS, et al. Lactylation signature identifies liver fibrosis phenotypes and traces fibrotic progression to hepatocellular carcinoma[J]. Front Immunol, 2024, 15: 1433393. DOI: 10.3389/fimmu.2024.1433393. [27] ZHAO TT, LI JF, ZHANG LT. Progress in the potential therapeutic mechanism of mesenchymal stem cell-derived exosomes for liver fibrosis[J]. Chin J Clin Pharmacol Ther, 2024, 29( 4): 475- 480. DOI: 10.12092/j.issn.1009-2501.2024.04.017.赵婷婷, 李俊峰, 张立婷. 间充质干细胞源性外泌体对肝纤维化潜在治疗机制的研究进展[J]. 中国临床药理学与治疗学, 2024, 29( 4): 475- 480. DOI: 10.12092/j.issn.1009-2501.2024.04.017. [28] TRIVEDI P, WANG S, FRIEDMAN SL. The power of plasticity-metabolic regulation of hepatic stellate cells[J]. Cell Metab, 2021, 33( 2): 242- 257. DOI: 10.1016/j.cmet.2020.10.026. [29] RHO H, TERRY AR, CHRONIS C, et al. Hexokinase 2-mediated gene expression via histone lactylation is required for hepatic stellate cell activation and liver fibrosis[J]. Cell Metab, 2023, 35( 8): 1406- 1423. DOI: 10.1016/j.cmet.2023.06.013. [30] ZHOU YQ, YAN JX, HUANG H, et al. The m6A reader IGF2BP2 regulates glycolytic metabolism and mediates histone lactylation to enhance hepatic stellate cell activation and liver fibrosis[J]. Cell Death Dis, 2024, 15( 3): 189. DOI: 10.1038/s41419-024-06509-9. [31] WU SJ, LI JH, ZHAN YF. H3K18 lactylation accelerates liver fibrosis progression through facilitating SOX9 transcription[J]. Exp Cell Res, 2024, 440( 2): 114135. DOI: 10.1016/j.yexcr.2024.114135. [32] TROGISCH FA, ABOUISSA A, KELES M, et al. Endothelial cells drive organ fibrosis in mice by inducing expression of the transcription factor SOX9[J]. Sci Transl Med, 2024, 16( 736): eabq4581. DOI: 10.1126/scitranslmed.abq4581. [33] YU YF, LI YN, ZHOU L, et al. Hepatic stellate cells promote hepatocellular carcinoma development by regulating histone lactylation: Novel insights from single-cell RNA sequencing and spatial transcriptomics analyses[J]. Cancer Lett, 2024, 604: 217243. DOI: 10.1016/j.canlet.2024.217243. [34] XIE N, TAN Z, BANERJEE S, et al. Glycolytic reprogramming in myofibroblast differentiation and lung fibrosis[J]. Am J Respir Crit Care Med, 2015, 192( 12): 1462- 1474. DOI: 10.1164/rccm.201504-0780OC. [35] DEWIDAR B, MEYER C, DOOLEY S, et al. TGF-β in hepatic stellate cell activation and liver fibrogenesis-updated 2019[J]. Cells, 2019, 8( 11): 1419. DOI: 10.3390/cells8111419. [36] GAO R, LI Y, XU Z, et al. Mitochondrial pyruvate carrier 1 regulates fatty acid synthase lactylation and mediates treatment of nonalcoholic fatty liver disease[J]. Hepatology, 2023, 78( 6): 1800- 1815. DOI: 10.1097/HEP.0000000000000279. [37] PAN RY, HE L, ZHANG J, et al. Positive feedback regulation of microglial glucose metabolism by histone H4 lysine 12 lactylation in Alzheimer’s disease[J]. Cell Metab, 2022, 34( 4): 634- 648 DOI: 10.1016/j.cmet.2022.02.013. [38] WANG PW, LIN TY, YANG PM, et al. Hepatic stellate cell modulates the immune microenvironment in the progression of hepatocellular carcinoma[J]. Int J Mol Sci, 2022, 23( 18): 10777. DOI: 10.3390/ijms231810777. [39] HUANG JL, FU YP, GAN W, et al. Hepatic stellate cells promote the progression of hepatocellular carcinoma through microRNA-1246-RORα-Wnt/β-Catenin axis[J]. Cancer Lett, 2020, 476: 140- 151. DOI: 10.1016/j.canlet.2020.02.012. [40] SUN C, XU WW, XIA YH, et al. PRDM16 from hepatic stellate cells-derived extracellular vesicles promotes hepatocellular carcinoma progression[J]. Am J Cancer Res, 2023, 13( 11): 5254- 5270. [41] YANG ZJ, YAN C, MA JQ, et al. Lactylome analysis suggests lactylation-dependent mechanisms of metabolic adaptation in hepatocellular carcinoma[J]. Nat Metab, 2023, 5( 1): 61- 79. DOI: 10.1038/s42255-022-00710-w. [42] WU QJ, LI X, LONG MH, et al. Integrated analysis of histone lysine lactylation(Kla)-specific genes suggests that NR6A1 OSBP2 and UNC119B are novel therapeutic targets for hepatocellular carcinoma[J]. Sci Rep, 2023, 13( 1): 18642. DOI: 10.1038/s41598-023-46057-4. [43] PAN LH, FENG F, WU JQ, et al. Demethylzeylasteral targets lactate by inhibiting histone lactylation to suppress the tumorigenicity of liver cancer stem cells[J]. Pharmacol Res, 2022, 181: 106270. DOI: 10.1016/j.phrs.2022.106270. [44] JIN J, BAI L, WANG DY, et al. SIRT3-dependent delactylation of cyclin E2 prevents hepatocellular carcinoma growth[J]. EMBO Rep, 2023, 24( 5): e56052. DOI: 10.15252/embr.202256052. -


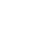 PDF下载 ( 1263 KB)
PDF下载 ( 1263 KB)
 下载:
下载:


