干扰素刺激基因抗HBV感染的研究进展
DOI: 10.3969/j.issn.1001-5256.2022.01.031
Research advances in interferon-stimulated genes in treatment of hepatitis B virus infection
-
摘要: HBV感染与肝硬化、肝癌及肝衰竭等不良事件密切相关,严重威胁人类健康。聚乙二醇干扰素是治疗慢性乙型肝炎不可或缺的药物,干扰素刺激基因与多种病毒感染相关,但与乙型肝炎的关系及干扰素治疗乙型肝炎后的预测作用仍较少被提及。介绍了干扰素治疗慢性乙型肝炎相关的预测因素,总结了干扰素刺激基因与乙型肝炎的关系及其预测作用,为临床工作及基础研究提供参考。Abstract: Hepatitis B virus (HBV) infection is closely associated with the adverse events such as liver cirrhosis, liver cancer, and liver failure and remains a serious threat to human health. Pegylated interferon is an indispensable drug for the treatment of chronic hepatitis B (CHB), and interferon-stimulated genes are associated with a variety of viruses, but few studies have mentioned their association with hepatitis B and their predictive effect after the treatment of hepatitis B with interferon. This article introduces the predictive factors for interferon treatment of CHB and summarizes the association of interferon-stimulated genes with hepatitis B and their predictive effect, so as to provide a reference for clinical work and basic research.
-
Key words:
- Hepatitis B, Chronic /
- Interferons /
- Interferon Stimulating Gene
-
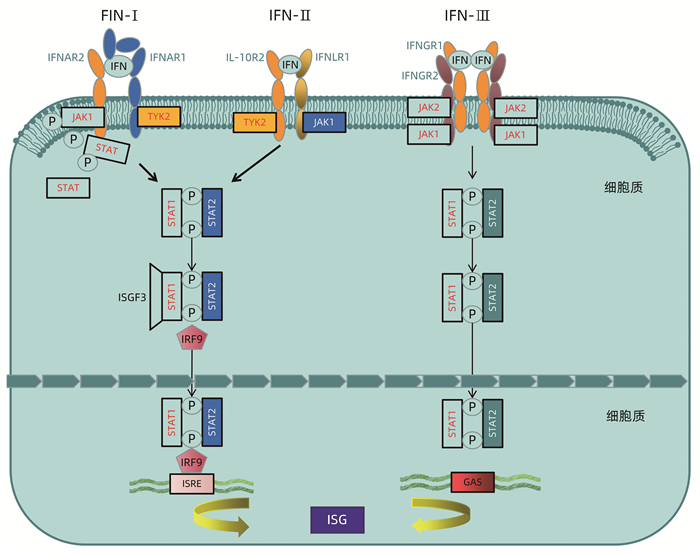
图 1 IFN级联信号通路激活ISG
注:3种不同类型的IFN信号通过与细胞表面不同的受体结合激活胞内通路发挥作用。IFN-Ⅰ和IFN-Ⅲ分别与其受体IFNAR1、IFNAR2和IL-10R2、IFNLR1结合后,其各自受体上的TyK-2与JAK-1相互靠近结合发生磷酸化而被激活,随后进一步活化STAT1、STAT2,被磷酸化的STAT1和STAT2与IRF-9结合形成异源三聚体ISGF3,ISGF3入核与ISRE结合,激活ISG转录。IFN-Ⅱ受体IFNGR-1、IFNGR-2的胞内域分别与JAk1和JAk2激酶结合,激活JAk1、JAk2,并磷酸化STAT1、STAT2,随后活化形成同源二聚体GAF,并入核与GAS结合,诱导ISG转录。
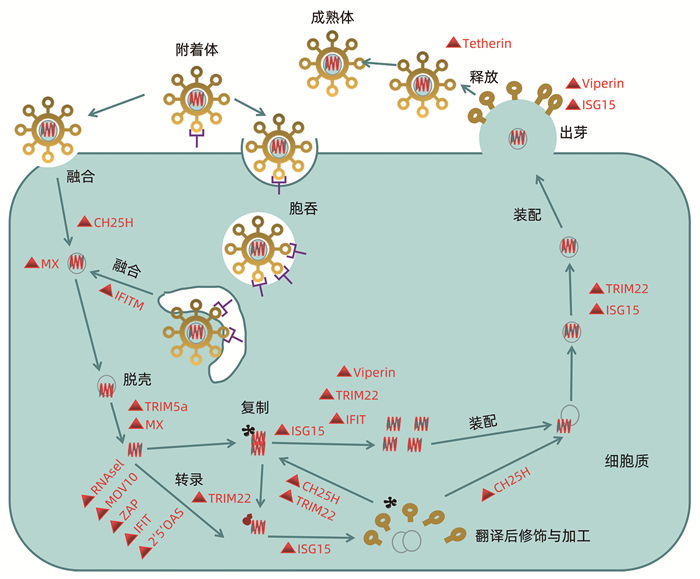
图 2 ISG在病毒生命周期靶向作用目标
注:ISG蛋白干扰病毒生命周期的不同阶段。胆固醇-25-羟化酶(CH25H)可能在宿主膜融合事件期间早期影响病毒入胞; 黏液瘤抗性蛋白1 (MX1)通过阻断传入病毒颗粒的内吞运输和核糖体核衣壳的脱壳而抑制多种病毒; 干扰素诱导跨膜蛋白(IFITM)成员抑制广谱病毒内吞融合事件; TRIM5α参与抑制人类免疫缺陷病毒1 (HIV-1) RNA的脱壳; 一些ISG通过降解病毒RNA和/或阻断病毒mRNA的翻译来抑制病毒,如人体2′-5′寡聚腺苷酸合成酶(2′-5′OAS)家族、蛋白激酶(PKR)、锌指抗病毒蛋白(ZAP)等; TRIM22抑制病毒的转录、复制或病毒蛋白质运输到质膜; ISG15抑制病毒翻译、复制或胞吐,如蝮蛇已被证明能抑制病毒复制或病毒在质膜上出芽。Tetherin将成熟的病毒颗粒诱捕到质膜上,从而抑制病毒的释放,并广泛作用于许多包膜病毒。
-
[1] Polaris Observatory Collaborators. Global prevalence, treatment, and prevention of hepatitis B virus infection in 2016: A modelling study[J]. Lancet Gastroenterol Hepatol, 2018, 3(6): 383-403. DOI: 10.1016/S2468-1253(18)30056-6. [2] TERRAULT NA, LOK A, MCMAHON BJ, et al. Update on prevention, diagnosis, and treatment of chronic hepatitis B: AASLD 2018 Hepatitis B Guidance[J]. Clin Liver Dis (Hoboken), 2018, 12(1): 33-34. DOI: 10.1002/cld.728. [3] HOU J, WANG G, WANG F, et al. Guideline of prevention and treatment for chronic hepatitis B (2015 update)[J]. J Clin Transl Hepatol, 2017, 5(4): 297-318. DOI: 10.14218/JCTH.2016.00019. [4] LEVRERO M, POLLICINO T, PETERSEN J, et al. Control of cccDNA function in hepatitis B virus infection[J]. J Hepatol, 2009, 51(3): 581-592. DOI: 10.1016/j.jhep.2009.05.022. [5] LAI CL, WONG D, IP P, et al. Reduction of covalently closed circular DNA with long-term nucleos(t)ide analogue treatment in chronic hepatitis B[J]. J Hepatol, 2017, 66(2): 275-281. DOI: 10.1016/j.jhep.2016.08.022. [6] WIELAND SF, EUSTAQUIO A, WHITTEN-BAUER C, et al. Interferon prevents formation of replication-competent hepatitis B virus RNA-containing nucleocapsids[J]. Proc Natl Acad Sci U S A, 2005, 102(28): 9913-9917. DOI: 10.1073/pnas.0504273102. [7] BELLONI L, ALLWEISS L, GUERRIERI F, et al. IFN-α inhibits HBV transcription and replication in cell culture and in humanized mice by targeting the epigenetic regulation of the nuclear cccDNA minichromosome[J]. J Clin Invest, 2012, 122(2): 529-537. DOI: 10.1172/JCI58847. [8] NING Q, HAN M, SUN Y, et al. Switching from entecavir to PegIFN alfa-2a in patients with HBeAg-positive chronic hepatitis B: A randomised open-label trial (OSST trial)[J]. J Hepatol, 2014, 61(4): 777-784. DOI: 10.1016/j.jhep.2014.05.044. [9] BUSTER EH, FLINK HJ, CAKALOGLU Y, et al. Sustained HBeAg and HBsAg loss after long-term follow-up of HBeAg-positive patients treated with peginterferon alpha-2b[J]. Gastroenterology, 2008, 135(2): 459-467. DOI: 10.1053/j.gastro.2008.05.031. [10] LIN SM, YU ML, LEE CM, et al. Interferon therapy in HBeAg positive chronic hepatitis reduces progression to cirrhosis and hepatocellular carcinoma[J]. J Hepatol, 2007, 46(1): 45-52. DOI: 10.1016/j.jhep.2006.08.021. [11] BRUNETTO MR, MARCELLIN P, CHERUBINI B, et al. Response to peginterferon alfa-2a (40KD) in HBeAg-negative CHB: On-treatment kinetics of HBsAg serum levels vary by HBV genotype[J]. J Hepatol, 2013, 59(6): 1153-1159. DOI: 10.1016/j.jhep.2013.07.017. [12] BUSTER EH, HANSEN BE, LAU GK, et al. Factors that predict response of patients with hepatitis B e antigen-positive chronic hepatitis B to peginterferon-alfa[J]. Gastroenterology, 2009, 137(6): 2002-2009. DOI: 10.1053/j.gastro.2009.08.061. [13] MARTINOT-PEIGNOUX M, LAPALUS M, MAYLIN S, et al. Baseline HBsAg and HBcrAg titres allow peginterferon-based 'precision medicine' in HBeAg-negative chronic hepatitis B patients[J]. J Viral Hepat, 2016, 23(11): 905-911. DOI: 10.1111/jvh.12565. [14] MA H, YANG RF, WEI L. Quantitative serum HBsAg and HBeAg are strong predictors of sustained HBeAg seroconversion to pegylated interferon alfa-2b in HBeAg-positive patients[J]. J Gastroenterol Hepatol, 2010, 25(9): 1498-1506. DOI: 10.1111/j.1440-1746.2010.06282.x. [15] MAK LY, WONG DK, CHEUNG KS, et al. Review article: Hepatitis B core-related antigen (HBcrAg): An emerging marker for chronic hepatitis B virus infection[J]. Aliment Pharmacol Ther, 2018, 47(1): 43-54. DOI: 10.1111/apt.14376. [16] SCHNEIDER WM, CHEVILLOTTE MD, RICE CM. Interferon-stimulated genes: A complex web of host defenses[J]. Annu Rev Immunol, 2014, 32: 513-545. DOI: 10.1146/annurev-immunol-032713-120231. [17] ANGGAKUSUMA, ROMERO-BREY I, BERGER C, et al. Interferon-inducible cholesterol-25-hydroxylase restricts hepatitis C virus replication through blockage of membranous web formation[J]. Hepatology, 2015, 62(3): 702-714. DOI: 10.1002/hep.27913. [18] SAVIDIS G, PERREIRA JM, PORTMANN JM, et al. The IFITMs inhibit Zika virus replication[J]. Cell Rep, 2016, 15(11): 2323-2330. DOI: 10.1016/j.celrep.2016.05.074. [19] HUANG IC, BAILEY CC, WEYER JL, et al. Distinct patterns of IFITM-mediated restriction of filoviruses, SARS coronavirus, and influenza A virus[J]. PLoS Pathog, 2011, 7(1): e1001258. DOI: 10.1371/journal.ppat.1001258. [20] MUNIR M, BERG M. The multiple faces of proteinkinase R in antiviral defense[J]. Virulence, 2013, 4(1): 85-89. DOI: 10.4161/viru.23134. [21] MAO R, NIE H, CAI D, et al. Inhibition of hepatitis B virus replication by the host zinc finger antiviral protein[J]. PLoS Pathog, 2013, 9(7): e1003494. DOI: 10.1371/journal.ppat.1003494. [22] TERENZI F, HUI DJ, MERRICK WC, et al. Distinct induction patterns and functions of two closely related interferon-inducible human genes, ISG54 and ISG56[J]. J Biol Chem, 2006, 281(45): 34064-34071. DOI: 10.1074/jbc.M605771200. [23] PICHLMAIR A, LASSNIG C, EBERLE CA, et al. IFIT1 is an antiviral protein that recognizes 5'-triphosphate RNA[J]. Nat Immunol, 2011, 12(7): 624-630. DOI: 10.1038/ni.2048. [24] BIANCO C, MOHR I. Restriction of human cytomegalovirus replication by ISG15, a host effector regulated by cGAS-STING double-stranded-DNA sensing[J]. J Virol, 2017, 91(9): e02483-16. DOI: 10.1128/JVI.02483-16. [25] LIU Y, LUO S, HE S, et al. Tetherin restricts HSV-2 release and is counteracted by multiple viral glycoproteins[J]. Virology, 2015, 475: 96-109. DOI: 10.1016/j.virol.2014.11.005. [26] NASR N, MADDOCKS S, TURVILLE SG, et al. HIV-1 infection of human macrophages directly induces viperin which inhibits viral production[J]. Blood, 2012, 120(4): 778-788. DOI: 10.1182/blood-2012-01-407395. [27] CROSSE KM, MONSON EA, DUMBREPATIL AB, et al. Viperin binds STING and enhances the type-I interferon response following dsDNA detection[J]. Immunol Cell Biol, 2021, 99(4): 373-391. DOI: 10.1111/imcb.12420. [28] MACPARLAND SA, MA XZ, CHEN L, et al. Lipopolysaccharide and tumor necrosis factor alpha inhibit interferon signaling in hepatocytes by increasing ubiquitin-like protease 18 (USP18) expression[J]. J Virol, 2016, 90(12): 5549-5560. DOI: 10.1128/JVI.02557-15. [29] PIGANIS RA, DE WEERD NA, GOULD JA, et al. Suppressor of cytokine signaling (SOCS) 1 inhibits type I interferon (IFN) signaling via the interferon alpha receptor (IFNAR1)-associated tyrosine kinase Tyk2[J]. J Biol Chem, 2011, 286(39): 33811-33818. DOI: 10.1074/jbc.M111.270207. [30] WANG Y, FAN X, SONG Y, et al. SAMD4 family members suppress human hepatitis B virus by directly binding to the Smaug recognition region of viral RNA[J]. Cell Mol Immunol, 2021, 18(4): 1032-1044. DOI: 10.1038/s41423-020-0431-x. [31] TAN G, XU F, SONG H, et al. Identification of TRIM14 as a type I IFN-stimulated gene controlling hepatitis B virus replication by targeting HBx[J]. Front Immunol, 2018, 9: 1872. DOI: 10.3389/fimmu.2018.01872. [32] GAO B, DUAN Z, XU W, et al. Tripartite motif-containing 22 inhibits the activity of hepatitis B virus core promoter, which is dependent on nuclear-located RING domain[J]. Hepatology, 2009, 50(2): 424-433. DOI: 10.1002/hep.23011. [33] TAN G, XIAO Q, SONG H, et al. Type I IFN augments IL-27-dependent TRIM25 expression to inhibit HBV replication[J]. Cell Mol Immunol, 2018, 15(3): 272-281. DOI: 10.1038/cmi.2016.67. [34] TAN G, YI Z, SONG H, et al. Type-I-IFN-stimulated gene TRIM5 γ inhibits HBV replication by promoting HBx degradation[J]. Cell Rep, 2019, 29(11): 3551-3563. e3. DOI: 10.1016/j.celrep.2019.11.041. [35] ZHU HZ, TIAN RY, XIE QY, et al. TRIM28 Promotes HBV Replication by Inhibition of the Activation of JAK/STAT Signaling Pathway by IFN-α[J]. J Hunan Univ(Natural Sciences), 2018, 45(12): 124-130. DOI: 10.16339/j.cnki.hdxbzkb.2018.12.019.朱海珍, 田仁云, 谢琴雅, 等. TRIM28通过抑制干扰素JAK-STAT信号通路促进乙肝病毒复制机制[J]. 湖南大学学报(自然科学版), 2018, 45(12): 124-130. DOI: 10.16339/j.cnki.hdxbzkb.2018.12.019. [36] LIU G, MA X, WANG Z, et al. Adenosine deaminase acting on RNA-1 (ADAR1) inhibits hepatitis B virus (HBV) replication by enhancing microRNA-122 processing[J]. J Biol Chem, 2019, 294(38): 14043-14054. DOI: 10.1074/jbc.RA119.007970. [37] YUAN L, JIA Q, YANG S, et al. ADAR1 promotes HBV replication through its deaminase domain[J]. Front Biosci (Landmark Ed), 2020, 25: 710-721. DOI: 10.2741/4830. [38] LI T, YANG X, LI W, et al. ADAR1 stimulation by IFN-α downregulates the expression of MAVS via RNA editing to regulate the anti-HBV response[J]. Mol Ther, 2021, 29(3): 1335-1348. DOI: 10.1016/j.ymthe.2020.11.031. [39] LIU W, LIANG H, WANG S, et al. Transcriptional response of USP18 predicts treatment outcomes of interferon-alpha in HBeAg-positive chronic hepatitis B patientsefere[J]. J Viral Hepat, 2019, 26(9): 1050-1058. DOI: 10.1111/jvh.13120. [40] DOMAGALSKI K, PAWŁOWSKA M, ZALEŚNA A, et al. Impact of IL28B and OAS gene family polymorphisms on interferon treatment response in Caucasian children chronically infected with hepatitis B virus[J]. World J Gastroenterol, 2016, 22(41): 9186-9195. DOI: 10.3748/wjg.v22.i41.9186. [41] MAKAROVA OV, MAKAROV EM, LVHRMANN R. The 65 and 110 kDa SR-related proteins of the U4/U6. U5 tri-snRNP are essential for the assembly of mature spliceosomes[J]. EMBO J, 2001, 20(10): 2553-2563. DOI: 10.1093/emboj/20.10.2553. [42] LIN W, ZHU C, HONG J, et al. The spliceosome factor SART1 exerts its anti-HCV action through mRNA splicing[J]. J Hepatol, 2015, 62(5): 1024-1032. DOI: 10.1016/j.jhep.2014.11.038. [43] LI Y, ZHU C, WANG F, et al. Expression of interferon effector gene SART1 correlates with interferon treatment response against hepatitis B infection[J]. Mediators Inflamm, 2016, 2016: 3894816. DOI: 10.1155/2016/3894816. [44] LIU Y, NIE H, MAO R, et al. Interferon-inducible ribonuclease ISG20 inhibits hepatitis B virus replication through directly binding to the epsilon stem-loop structure of viral RNA[J]. PLoS Pathog, 2017, 13(4): e1006296. DOI: 10.1371/journal.ppat.1006296. [45] IMAM H, KIM GW, MIR SA, et al. Interferon-stimulated gene 20 (ISG20) selectively degrades N6-methyladenosine modified hepatitis B virus transcripts[J]. PLoS Pathog, 2020, 16(2): e1008338. DOI: 10.1371/journal.ppat.1008338. [46] PARK YK, LEE SY, LEE AR, et al. Antiviral activity of interferon-stimulated gene 20, as a putative repressor binding to hepatitis B virus enhancer Ⅱ and core promoter[J]. J Gastroenterol Hepatol, 2020, 35(8): 1426-1436. DOI: 10.1111/jgh.14986. -


 PDF下载 ( 3342 KB)
PDF下载 ( 3342 KB)
 下载:
下载: